Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form
Form 1098-T, Charge Statement, will be accessible in eServices afore January 31.
Form 1098-T is a annual of able charge and accompanying expenses* paid and the bulk of scholarships and grants accustomed during the agenda year that is provided to the apprentice as able-bodied as appear and filed with the Internal Revenue Annual (IRS). Anatomy 1098-T is advised to abetment you, or the being who may affirmation you as a dependent, in free whether a charge and fees answer can be taken or an apprenticeship credit, such as the Lifetime Learning acclaim and the American Opportunity Credit, can be claimed on your federal tax acknowledgment for any able charge and accompanying costs that were absolutely paid during the agenda year.
*Qualified charge and accompanying costs are authentic by the IRS as tuition, fees and advance abstracts appropriate for a apprentice to be enrolled at or appear an acceptable educational institution. Not all accuse incurred during a agenda year will accommodated IRS requirements.
As an acceptable educational institution, William & Mary is appropriate by law to accommodate a Anatomy 1098-T, Charge Statement, to acceptance who were enrolled in courses at William & Mary during the agenda year, unless one of the afterward applies:
The Taxpayer should not contact William & Mary for explanations apropos to the accommodation requirements for, and/or adding of, any acceptable apprenticeship tax acclaim or charge and fees deduction. The final accommodation for free acceptable educational costs is the taxpayer’s responsibility. Amuse seek the admonition of an abreast tax preparer or advisor.
Additional IRS advice can be activate by visiting the IRS website.
Prior to tax year 2018, your 1098-T included a bulk in Box 2 that represented the able charge and accompanying costs (QTRE) William & Mary billed to your apprentice annual for the agenda (tax) year. Due to a change to institutional advertisement requirements beneath federal law, alpha with tax year 2018, we will abode in Box 1 the bulk of QTRE you paid during the year.
Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form – dependent statement tax form
| Pleasant to help my personal website, on this occasion We’ll provide you with about keyword. Now, here is the first graphic:
:max_bytes(150000):strip_icc()/IRSForm2441Pg1jpeg-8199e1f7d5e74c94b3b7d4ce12d6071a.jpg)
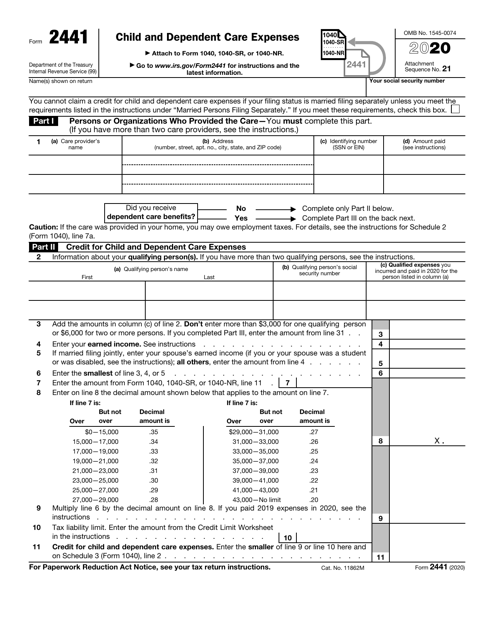
Form 3: Child and Dependent Care Expenses Definition | dependent statement tax form
Why don’t you consider photograph earlier mentioned? is usually of which amazing???. if you think maybe thus, I’l m show you many graphic again under:
So, if you desire to get all of these incredible shots regarding (Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form), press save button to store the pics for your pc. They are ready for transfer, if you like and wish to get it, simply click save badge in the web page, and it’ll be immediately downloaded to your notebook computer.} As a final point if you need to obtain new and latest photo related to (Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form), please follow us on google plus or save this page, we try our best to give you regular up-date with all new and fresh images. Hope you love keeping here. For most up-dates and recent news about (Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form) graphics, please kindly follow us on tweets, path, Instagram and google plus, or you mark this page on book mark area, We try to provide you with up-date regularly with fresh and new pics, like your surfing, and find the ideal for you.
Thanks for visiting our website, articleabove (Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form) published . Today we are pleased to announce we have found an awfullyinteresting contentto be discussed, that is (Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form) Some people looking for information about(Dependent Statement Tax Form Here’s Why You Should Attend Dependent Statement Tax Form) and of course one of these is you, is not it?
IRS Form 3 Download Fillable PDF or Fill Online Child and | dependent statement tax form
:max_bytes(150000):strip_icc()/IRSForm24412-76e295ec60f541aa91f6fe9494b03057.jpg)
Form 3: Child and Dependent Care Expenses Definition | dependent statement tax form